The Record Is Complete. The Picture Is Wrong.
A patient record can be comprehensive and clinically wrong at the same time. The AI reasons from what it receives. The failure is upstream, in the data, not in the model. That is the hallucination type nobody is naming.
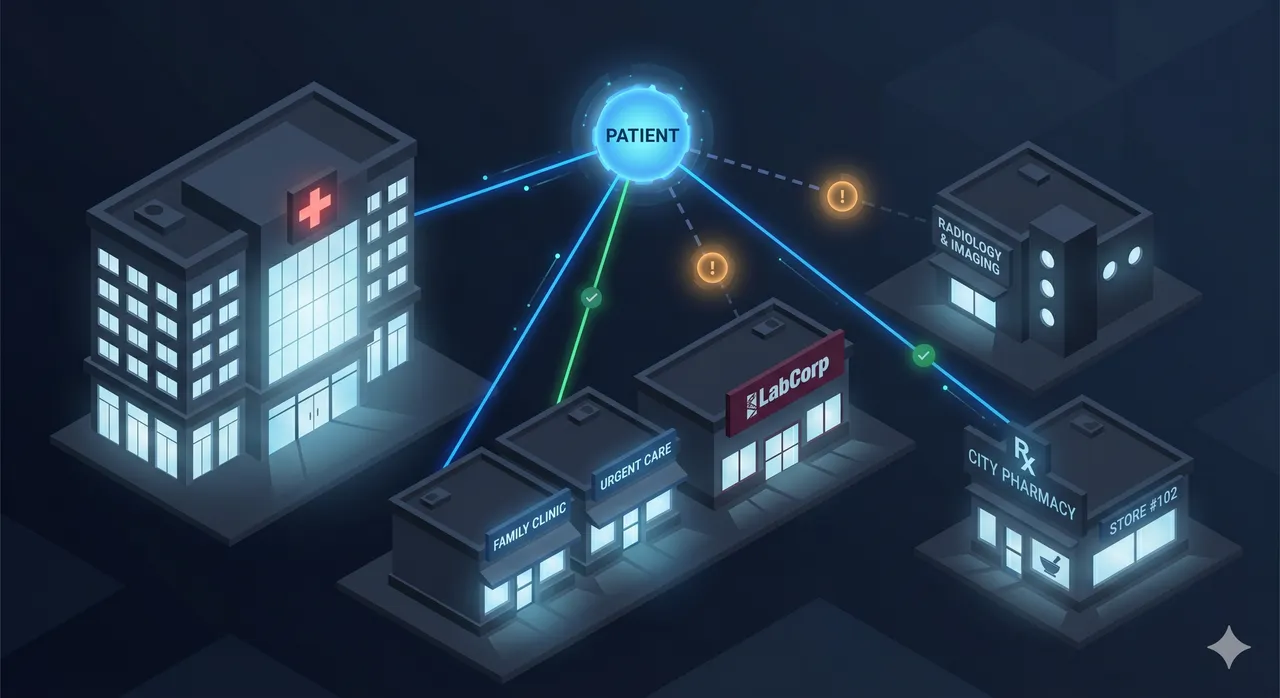
Last night Amazon AI filled a prescription for me in two hours. The last time I tried the same thing through my regular provider it took a week, a scheduled visit, and a co-pay that cost more than the medication.
The medication is an antibiotic cream, sold over the counter at any European pharmacy for a few euros. In the US it requires a prescription. I have never fully understood why.
I found it while searching on Amazon, which now has detailed pharmaceutical pages for prescription drugs. I clicked the request button and entered the funnel. Amazon OneCare. Pay per visit at $29, waived with some introductory Prime offer I did not fully understand. The AI questionnaire took five minutes. Two hours later a human provider had reviewed my submission and sent the prescription to my local pharmacy. I chose a pharmacy two blocks from my house. Nobody pushed back.
But before the questionnaire started, something else happened. The system surfaced a list of medications and health conditions pulled from my connected health records. Years of encounters across multiple providers, assembled in one place. It was extensive. It asked me to confirm which items were still relevant.
I looked at the list for a while.
There were conditions I did not recognize. Events from more than a decade ago, clinically resolved long before any of these platforms existed. There were medications prescribed once for acute problems that had cleared completely. Allergies that had been formally evaluated and removed from my chart at one institution, still listed as active because the correction had never traveled to the connected systems. The record was comprehensive. It was also, in any clinically meaningful sense, wrong. Dangerously wrong.
This is the problem nobody is talking about.
The platforms building the next generation of consumer health AI, including ChatGPT Health, Amazon Health, Apple Health, and Microsoft Copilot Health, have made a bet that the path to better health outcomes runs through data aggregation. Give patients control over their own records. Let them connect providers. Assemble a unified view. Then let AI reason from the complete picture.
The premise is correct. The execution has a structural flaw that will not resolve on its own.
Patient-mediated aggregation creates two simultaneous failure modes that pull in opposite directions. The first is the gap you cannot see. The second is the noise you cannot filter. They coexist in every aggregated record, and current systems are designed to handle neither.
The gap you cannot see
The promise of connecting patient records implies that connection equals completeness. It does not. Health information is generated across dozens of systems: primary care EHRs, specialist platforms, pharmacy databases, insurance claims, imaging systems, lab networks, wearables. Each speaks a different data language. No single consent flow reaches all of them.
The consequences are not theoretical. Research shows that only 16% of patients with lab-confirmed chronic kidney disease have CKD documented on their problem list. Meaning 84% of that diagnosis is clinically active and completely invisible to any aggregation platform. Studies on EHR problem list completeness for common chronic conditions find that between 20% and 40% of known diagnoses are simply absent, not because the conditions do not exist, but because documentation did not follow the patient across systems.
The AI observes the connected record and reasons from what it sees. The gap is not marked as a gap. It appears as the absence of clinical activity, which the system may interpret as normal health status. A patient with unconnected monitoring from a specialist clinic looks, in the aggregated view, like a patient who does not need monitoring. The output is a hallucination. Not because the model invented anything, but because the premise was incomplete.
The noise you cannot filter
The second failure runs in the opposite direction. Clinical records are primarily additive. Diagnoses accumulate. Prescriptions stack. Allergies are added. Very little is ever formally removed or marked resolved in a way that travels across systems.
FHIR, the standard governing how most of this data moves, has the right architecture. The Condition resource has a clinicalStatus field with values including active, inactive, remission, and resolved. The MedicationRequest has a status field. The AllergyIntolerance resource can be marked inactive or refuted. The specification is sound.
The implementation is not. Research examining real-world FHIR data finds that these status fields are inconsistently authored at the source, often defaulted to active during EHR entry because updating them takes time nobody has, and frequently overwritten during FHIR export with whatever the mapping layer assumes when the field is ambiguous. A condition entered as active in 2015 and never updated arrives in your aggregated record in 2026 as active. The FHIR pipeline did its job correctly. The information is wrong.
The medication accuracy problem is severe. Studies at hospital admission find that almost half of medication discrepancies come from failure to capture what the patient was actually taking, 85% of medication errors originate in medication histories, and over 40% of medication errors in hospitals trace back specifically to poor reconciliation at care transitions. An estimated 770,000 Americans are injured or die annually from adverse drug events tied to incomplete or inaccurate medication data. The allergy picture is similarly compromised: more than 90% of penicillin allergy labels are inaccurate when formally evaluated, yet 31% of patients who have been formally de-labeled at a hospital still carry an active penicillin allergy at their pharmacy, because the correction did not propagate across systems.
When AI reasons from this aggregated view, it is not reasoning from a complete picture. It is reasoning from a comprehensive historical archive that has been stripped of the clinical annotations that distinguish then from now. The output is also a hallucination. A different kind: the premise is polluted rather than absent, but the result is the same confident wrong answer grounded in data the AI had no reason to question.
This is the part that tends to get lost in discussions about AI hallucinations. We have built an entire industry response around the idea that hallucinations are random confabulations, facts the model invented with no grounding. The prescribed remedy is the human in the loop: a reviewer who checks the AI's output, traces it back to a source, and flags what does not check out.
That defense works for fabrications. It does not work for this.
I wrote previously about a taxonomy of AI hallucination types developed by researchers at MIT, Harvard Medical School, and Google Research. The taxonomy maps five structured failure modes: factual errors, outdated references, spurious correlations, fabricated sources, and incomplete reasoning chains. Every cluster in that taxonomy is a model-layer failure. The model did something wrong with the data it received.
The failure mode described in this article is not in that taxonomy. It belongs upstream of all five clusters. The model does nothing wrong. It reasons correctly from the data it receives. The data is wrong. The output is confidently, specifically, and traceable-to-a-real-source wrong. When a physician or a patient reviews an AI recommendation and sees it citing a real record, a real allergy entry, a real diagnosis code, a real medication history, they are not looking at a hallucination in any sense they have been trained to recognize. They are looking at a grounded, well-reasoned output from a flawed premise. The human reviewer approves it because it checks out. It does not check out.
This is the sixth cluster the taxonomy did not cover: data-layer hallucination. The model is working correctly. The input is wrong. The wrongness is perfectly explainable after something goes wrong and almost invisible before it does. The AI did not make anything up. It read the record faithfully and reasoned from it precisely. The record was wrong, and that wrongness was invisible to every layer of the system designed to catch errors.
The mechanism: what actually happens when data moves
Most of these problems trace to a single point of failure. Clinical records are authored and maintained by humans working under time pressure inside a specific institution's workflow. The problem list is owned by whoever last touched it. The medication list is updated when a pharmacist reconciles at admission, but not consistently between admissions. The allergy record is corrected by the allergist, but the correction lives in the allergist's system and may never reach the primary care EHR.
FHIR was designed to move data, not to verify it. It transfers what the source system provides. If the source system has an outdated status, FHIR faithfully reproduces the outdated status. If the field is missing, FHIR mapping layers frequently default to active because that is the safest assumption for a system that does not know what it does not know.
The aggregation platform receives this data and, under current architectures, cannot distinguish between an active condition and a condition that was never updated. The two look identical in the transferred record.
What a smarter aggregation layer would do
The research community has begun to address this using knowledge graph architectures. In a graph representation, every patient node is connected to condition nodes through explicitly typed edges. A patient with a diabetes node but no HbA1c observation edge is not a normal state. It is a detectable structural absence, a gap that is computable rather than invisible. Systems like GraphCare, developed at ICLR 2024 through a collaboration between the University of Illinois and GE HealthCare, apply this logic to clinical prediction and achieve substantial improvements in accuracy over flat-record approaches.
This logic has not yet been applied to data acquisition. It should be. If the system knows a patient has Type 2 diabetes, it already knows what a clinically complete record for that patient should contain. Regular HbA1c values. A current kidney function panel. An active medication from the standard diabetes pharmacopeia, or documentation of why none was prescribed. Annual retinal imaging. Blood pressure trends. Ideally a note from a dietitian. When any of these elements are absent, the system can flag the gap as a gap, not as normal absence, and use it to identify where additional data sources should be sought.
This is Condition-Aware Consolidation: using the clinical knowledge encoded in a patient's known conditions to generate an expected data footprint, measure the actual record against it, and treat discrepancies as signals rather than as nothing. The expectation set must be personalized, not static. A 75-year-old with Type 2 diabetes and a 30-year-old with gestational diabetes share a diagnosis code but require entirely different data footprints to be clinically meaningful.
Any data point can serve as the starting anchor. A confirmed diagnosis is the most reliable. A prescribed medication or a lab result can expand the search radius: the presence of metformin suggests looking for HbA1c and kidney function data; a retinal scan suggests looking for the ophthalmology system it originated from. But this directional logic must be handled carefully.
Using a data point to expand the search is not the same as using it to infer a diagnosis. Labetalol can treat high blood pressure or anxiety. Metformin is prescribed for pre-diabetes, polycystic ovarian syndrome, weight management, and off-label purposes. In PCOS populations alone, inferring Type 2 diabetes from a metformin prescription would be clinically wrong the majority of the time. An elevated HbA1c is a signal, not a conclusion. The moment the aggregation layer says this patient probably has Type 2 diabetes based on a medication, it has crossed from data engineering into clinical reasoning, and it has no authority to do that. The principle is expansion without inference: the reverse knowledge graph expands what the system looks for; it does not conclude what the patient has.
This boundary matters for regulatory as well as clinical reasons. The 2026 FDA guidance on AI-enabled clinical decision support draws a clear line between tools that support clinical judgment and tools that substitute for it. The former may be excluded from device regulation. The latter requires premarket review. A consolidation layer that generates an expected data footprint and flags gaps is supporting the clinician or the patient's verification step. A consolidation layer that generates a provisional diagnosis from medication patterns has entered regulated territory without a license.
A note on verification design and the source system objection
Two implementation challenges deserve naming. The first: asking patients to verify historical clinical data is not as simple as asking them to confirm a shipping address. Studies show patient engagement with health portal requests drops sharply when the prompt requires clinical judgment rather than simple confirmation. Any verification step must be designed as low cognitive load, using visual timelines, plain language, and clear defaults, not clinical prompts that most patients are not equipped to answer.
The second objection experts raise is the obvious one: why not simply clean the source systems? The answer is that ideal prescribing and ideal documentation practices would prevent most of this. They are also ten or more years away from being standard. Client-layer mitigation, building intelligence into the aggregation layer itself, is not a workaround. It is the near-term path to patient safety.
The gap in the current landscape
A review of the platforms currently operating in this space finds that none of them, including ChatGPT Health, Amazon Health, Apple Health, and Anthropic's Claude for Healthcare, currently employ conditional logic for data completeness. They focus on summarizing whatever is connected, with privacy safeguards and user disclaimers, but do not yet apply logic like: you have diabetes, here are the data elements commonly associated with that condition, and here is what appears to be missing from your connected record. Patient-initiated connection and manual review remain the only mechanisms for addressing gaps. The research community has demonstrated feasibility. The commercial implementation does not yet exist.
What this means for product designers
The platforms building consumer health AI today are solving the access problem. Amazon's two-hour prescription delivery is genuinely impressive. ChatGPT Health's natural language interface to connected records removes friction that has frustrated patients for years. The access and convenience improvements are real.
The data quality problem is the next layer. It will surface as AI outputs that seem authoritative but are based on records that are simultaneously incomplete and contaminated. The Washington Post tested ChatGPT Health and received an "F" health grade based on ten years of Apple Watch data. The reporter's physician and cardiologist Eric Topol independently called the grade baseless. The system lacked clinical context that would have changed the interpretation entirely.
The solution is not to wait for cleaner data. Healthcare data will never be uniformly clean. The solution is to build an aggregation layer that understands what it should be looking for, flags what is missing, and does not treat historical artifacts as current clinical facts.
Aggregating health data is not the same as understanding it. The platforms that figure out the difference will not just deliver access. They will deliver something more valuable: a clinical picture the AI can actually reason from.
This article is part of an ongoing series on healthcare data, AI infrastructure, and the systems that connect them.